Apenas para uso em pesquisa. Não utilizar em procedimentos diagnósticos.
On November 24, 2021, variant B.1.1.529 was first identified in South Africa, and just two days later, the WHO announced that B.1.1.529 constituted a Variant of Concern (VOC), named “Omicron.”
There are more than 30 mutations in the S gene of the B.1.1.529 variant. Omicron has a higher binding affinity than the original SARS-CoV-2, showing levels comparable to those observed with the Delta variant. Some of these mutations have been previously detected in earlier variants, such as Alpha and Delta, and are associated with increased transmissibility and immune evasion.
Many other identified mutations are not yet fully characterized and have not been found in other circulating variants. Investigations continue to be conducted to determine the impact of these changes. The Omicron family includes the Pango B.1.1.529 lineage and the descendant lineages BA.1, BA.1.1, BA.2, and BA.3.
Precise: Identification of the Omicron variant through simultaneous detection of ORF1ab and four specific S-gene mutations.
Smart: Automatic interpretation of results on STC-96A and STC-96A Plus* systems.
Versatile: Semi-automatic tool available for other real-time PCR instruments
Reliable: Endogenous internal control (IC) ensures reliability throughout the process
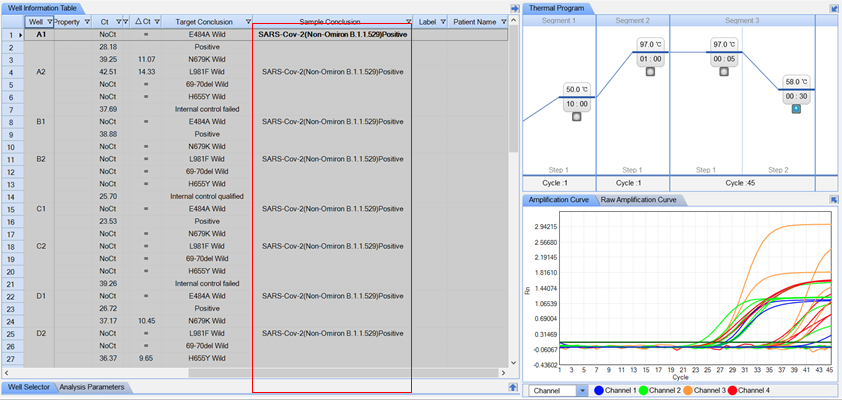
| Targets: |
ORF1ab and S-gene mutations E484A, N679K, L981F, and H655Y |
| Positive Percent Agreement | 100% |
| Negative Percent Agreement | 100% |
| Limit of Detection (LoD) | 500 copies/mL |
| Sample types |
Nasopharyngeal swab, oropharyngeal swab, and sputum |
Apenas para uso em pesquisa. Não utilizar em procedimentos diagnósticos.